DataJoint Enables Seamless Migration of CWL Pipelines to Its Governed, Reproducible Scientific Data Infrastructure
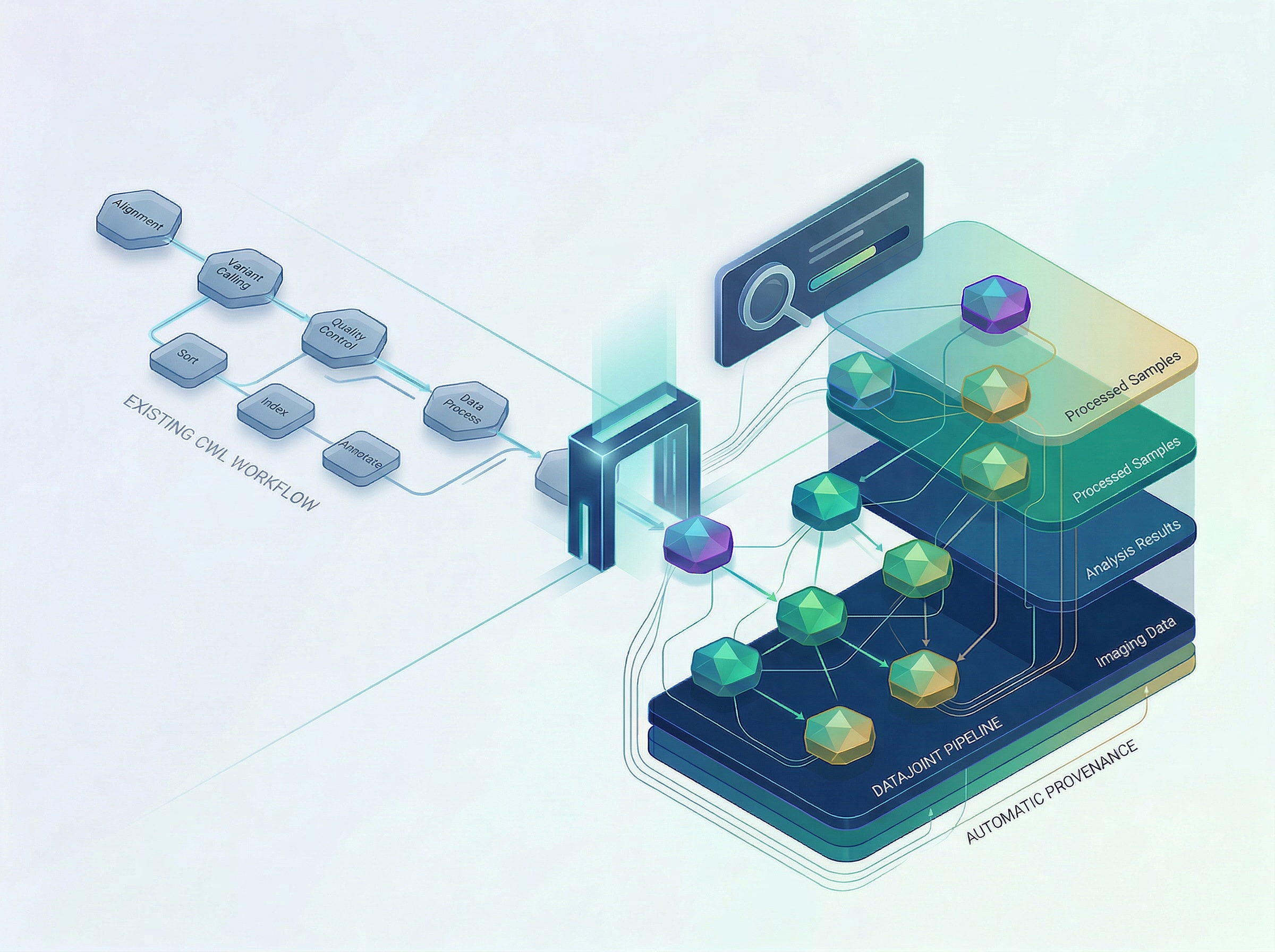
DataJoint today announced native support for converting Common Workflow Language (CWL) pipelines into DataJoint pipelines, enabling research organizations to immediately modernize existing scientific workflows — without sacrificing prior investment or starting from scratch.
CWL: Widely Adopted, But Increasingly Constrained
Common Workflow Language has become a de facto standard across pharmaceutical R&D, genomics, and academic research for defining portable, reproducible computational workflows. Major cloud and bioinformatics platforms support CWL natively, and it is broadly adopted across federally funded genomics programs and industry R&D consortia — making it one of the most widely deployed workflow standards in life sciences.
Yet CWL has recognized limitations in production environments: limited error handling and debugging, no native provenance tracking, poor support for partial re-runs when a step fails mid-pipeline, and no mechanism to query workflow state. As AI-driven research demands tighter auditability and reproducibility, these gaps create real scientific and operational risk.
What DataJoint Provides
DataJoint's CWL conversion layer reads existing CWL workflow definitions and executes them as native DataJoint pipelines. Research teams can extend these pipelines — mixing CWL definitions with DataJoint's Python-based schema framework — and run them in interpreted mode today, with compiled execution on the roadmap. Key capabilities include:
Automatic provenance. Every CWL step is backed by DataJoint's schema-driven provenance layer, creating a complete, queryable record of inputs, outputs, and computational history.
Granular retry and resilience. Failed steps can be individually retried or corrected without re-running the entire pipeline — a critical capability for long-running, high-cost workflows.
Queryable state. Workflow state is accessible via DataJoint's standard query syntax, enabling real-time monitoring and downstream analysis.
Natural parallelization. Pipelines are decomposed into discrete, independently executable steps that support cluster-level parallelism and graceful pause/resume without lost progress.
Structured entity database. Critically, DataJoint does not simply execute CWL workflows — it builds a structured database around the scientific entities those workflows produce. The conversion process involves explicitly defining the entities created at each stage (such as processed samples, imaging results, or analysis outputs) and the dependencies between them. This transforms a pipeline from a sequence of compute steps into a living, queryable scientific record — one that captures not just what ran, but what was produced, how it relates to other data, and how it can be reused.
"Scientific AI will only be as trustworthy as the data foundation beneath it. CWL gave the research community a powerful way to define workflows — DataJoint gives those workflows the provenance, traceability, and governance they need to support defensible science and AI-ready research at scale." — Jim Olson, CEO, DataJoint
Related posts
DataJoint Appoints Former Flywheel Exec Jim Olson as New CEO
A New Operating System for Science
Preprint: The DataJoint Model
Updates Delivered *Straight to Your Inbox*
Join the mailing list for industry insights, company news, and product updates delivered monthly.

.svg)






